
MetaDome was recently recognized as one of the most accessed articles of 2018-2019 in Human Mutation.
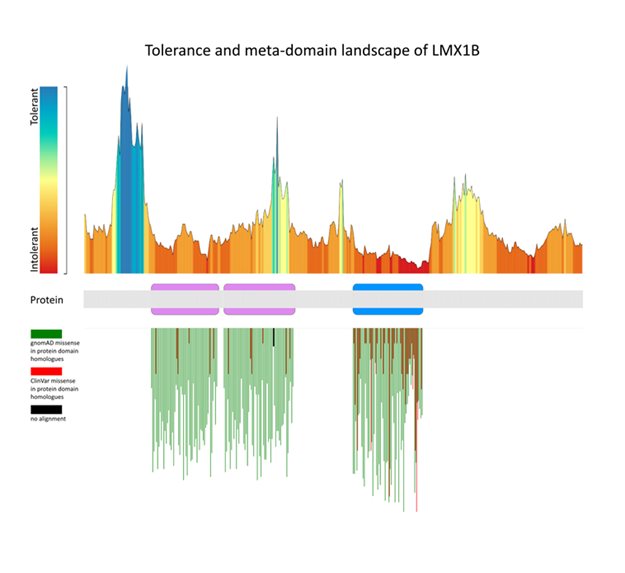
Researchers of the Radboudumc have developed MetaDome to survey regions in proteins that tolerate genetic variation. This helps to address the pathogenicity of genetic variants of unknown clinical significance. Using evolutionary relationships between protein domain homologues, MetaDome aggregates variation onto “meta-domains”. These meta-domains make the genetic variation map richer than individual protein domains. MetaDome additionally visualizes variation into a tolerance landscape, indicating regions within protein-coding genes that are more, or less, tolerant to variation.
The MetaDome was a project developed by Laurens van de Wiel, Christian Gilissen, and Gert Vriend and was first available online in spring 2018. Since then, it has been used by nearly 2,500 colleagues from various institutes across 64 countries and can be accessed here.
Related news items

Renee Salz wins prize for best presentation
10 May 2022 Renee Salz won the award for best oral presentation during the Rolduc genetics retreat, organized by the Dutch and Belgian Societies of Human Genetics in Brugge. Her presentation was titled "Variant effect prediction based on long read transcriptomes". go to page
Large NWA ORC grant awarded for national skin research: Next Generation ImmunoDermatology
23 March 2022Research for better treatment methods for chronic skin diseases.
go to page
Metakids-UMD grant for Purva Kulkarni
18 June 2020 The Metakids foundation has awarded 220.575 Euro to support the establishment of a multi-omics biomarker platform within the United for Metabolic Disease (UMD) consortium. go to page
Front cover Human Mutation
21 August 2019The MetaDome web server build to interpret genetic variants based on genetic tolerance and homologous protein domains is featured on the Cover of Human Mutation. MetaDome was developed by Laurens van de Wiel, Coos Baakman, Daan Gilissen, Joris Veltman, Gert Vriend and Christian Gilissen,
go to page
ZonMw grant to promote large scale (re)use of DNA data
22 November 2018 Collaborative effort towards enhanced reusability of DNA data facilitates future personalized medicine approaches. go to page
Publication in Human Mutation Using homologous relationships of protein domains in the human genome to interpret genetic variation
4 September 2017This publication is part of the PhD project of Laurens van de Wiel and shows how homologous protein domain relations may be used to interpret normal and pathogenic variation at a much finer scale than previously possible.
go to page